The Shannon Capacity of a Graph, 2
Posted by Tom Leinster
Graph theorists have many measures of how ‘big’ a graph is: order, size, diameter, radius, circumference, girth, chromatic number, and so on. Last time I told you about two of them: independence number and Shannon capacity. This time I’ll bring in two other measures of size that I’ve written about previously in contexts other than graph theory: maximum diversity and magnitude.
The post will end not with a bang, but a whimper. Although there’s something simple and clean to say about the connection between maximum diversity and standard graph invariants, the relationship with magnitude is much more mysterious, and I don’t have much more to say than ‘what’s going on?’. So there will be no satisfying conclusion. Consider yourself warned!
A crash course in diversity measurement Imagine an ecosystem consisting of a number of species. We know what proportions the various species are present in, and we know how similar the species are to one another. How can we quantify, in a single number, the ‘diversity’ of the ecosystem?
Over the years, many people have answered this question in many different ways. The answer I’m about to explain is the one given in a paper of mine with Christina Cobbold, called Measuring diversity: the importance of species similarity. I also posted about it here and on John’s blog. But I won’t assume you’ve read any of that.
Let’s say we have species, in respective proportions . Thus, is a probability vector: a finite sequence of nonnegative reals summing to . And let’s say that the similarity of one species to another is measured on a scale of to , with meaning ‘completely different’ and meaning ‘identical’. These similarities are the entries of an matrix , the similarity matrix, which I’ll assume is symmetric and has s down the diagonal (because any species is identical to itself).
The expected similarity between an individual from the th species and a randomly-chosen individual is . This measures how ‘ordinary’ the th species is. We can express this formula more neatly if we regard as a column vector: then the ordinariness of the th species is simply , the th entry of the column vector .
A community in which most species are very ordinary is not very diverse; conversely, a community in which most species are highly distinctive is very diverse. So the average ordinariness, across all species, measures the ecosystem’s lack of diversity. The average ordinariness is
To get a measure of diversity itself, we take the reciprocal:
What’s that doing on the left-hand side? Well, there’s a certain -ness to the right-hand side, since the denominator is the quadratic form . And in fact, this diversity measure is just one of a one-parameter family of measures.
The other members of the family come about by varying our interpretation of ‘average’ in the phrase ‘average ordinariness’. Just now, we interpreted it as ‘arithmetic mean’. But the arithmetic mean is only one member of a larger family: the power means. Let . The power mean of order of positive reals , weighted by a probability vector , is
In other words: transform each by taking the th power, form the arithmetic mean, then apply the inverse transformation. Clearly the formula doesn’t make sense for and , but we can fill in those gaps by taking the limit as or . This gives
In this notation, the formula for diversity given above is
Generally, for any , the diversity of order of the ecosystem is
This turns out to be a quantity between and , and can be thought of as the ‘effective number of species’. It takes its minimum value of when only one species is actually present, that is, for some . It takes its maximum value of when the species are all completely dissimilar (that is, is the identity matrix) and they’re present in equal proportions.
Different values of attach different amounts of significance to rare species, in a way that I won’t explain. But it’s important to recognize that the diversities of different orders really do measure different things. For example, it often happens that ecosystem A has greater diversity of order than ecosystem B, but less diversity of order .
If you know some information theory, you can recognize among these diversity measures some classical measures of entropy. For example, when (different species are deemed to have nothing whatsoever in common), we get the exponential of Rényi entropy:
The even more special case , gives the exponential of Shannon entropy.
Maximum diversity Now suppose we have a list of species of known similarities, and also suppose that we have control over their proportions or frequencies. In other words, we’ve fixed and and we can vary . How can we choose to maximize the diversity? And what is the value of that maximum diversity?
Well, these questions don’t entirely make sense. Remember that we have not just one diversity measure, but a one-parameter family of them. Perhaps the that maximizes diversity of order comes nowhere near maximizing diversity of order . And perhaps , the maximum diversity of order , is different from . After all, different values of give genuinely different measures.
So it comes as a surprise that, in fact, there’s a ‘best of all possible worlds’: a probability vector that maximizes diversity in all these uncountably many senses at once. And it’s a double surprise: all these different diversity measures have the same maximum value. In other words:
Theorem Let and let be an similarity matrix. Then:
There is a probability vector such that for all , we have ;
is independent of .
In particular, associated to any similarity matrix is a real number, its maximum diversity, defined as
for any . Which makes no difference: that’s the second part of the theorem.
How graphs come in Now consider the special case where all inter-species similarities are either or : each pair of species is deemed to be either totally dissimilar or totally identical. The matrix of similarities is then simply a symmetric matrix of s and s, with s down the diagonal.
A matrix with these properties is the same thing as a finite, reflexive, undirected graph — which in these posts I’m just referring to as graphs. That’s because a graph can be seen in terms of its adjacency matrix .
So, given a graph , we can ask: what is the maximum diversity of the matrix ? The official notation for this is , but I’ll write it as instead, and call it the maximum diversity of . What is it, explicitly?
It turns out that it’s exactly the independence number of . In Part 1 I mentioned that a set of vertices in a graph is said to be independent if no two elements have an edge between them, and that the independence number is the largest cardinality of an independent set in . So, the result is that
for all graphs . Or in words:
Maximum diversity is independence number.
There are uncountably many proofs. Because is equal to for any , you can choose whichever is most convenient and maximize the diversity of that order. If you take , it becomes pretty simple — I reckon that’s the easiest proof. (In Example 4.2 of my paper, I did it with , but that was probably less efficient.)
Back to Shannon capacity Last time, I gave you the definition of the Shannon capacity of a graph : it’s
where is the ‘strong product’ of graphs: the product in the category of (reflexive, as always) graphs. If we want, we can write the definition of with in place of , since they’re the same thing.
I mentioned that computing Shannon capacity is really hard: no one even knows the Shannon capacity of the 7-cycle! So people have put a lot of effort into finding quantities that are similar to , but easier to compute.
For example, suppose we can find a tight but easy-to-compute upper bound on . We already have plenty of lower bounds, namely, for any we like. So we’d then have sandwiched between two easy-to-compute bounds. For some graphs, the upper and lower bounds might even be equal, giving us the exact value of .
The best-known substitute for the Shannon capacity is called the the Lovász number, . I confess, I don’t have any feel for the definition (which we won’t need here), but here are some of its good properties:
It’s much more efficient to compute than the Shannon capacity.
It’s an upper bound: . (Yup, the quantity denoted by a small letter is bigger than the quantity denoted by a big letter.)
It’s multiplicative: . This formula helps us to compute the Lovász number of any graph of the form . Neither the independence number nor the Shannon capacity is multiplicative.
As Omar Antolín Camarena pointed out last time, even the Shannon capacity of the 5-cycle was unknown until Lovász defined his number. Here’s how Lovász did it. Having proved that is an upper bound for , he then showed that . But
and we calculated last time that . Hence can only be .
(There’s at least one other useful upper bound for , defined by Haemers and pointed out by Tobias last time, but I won’t say anything about it.)
Maximum diversity and magnitude We do know one nice quantity to which maximum diversity is closely related, and that’s magnitude.
I’ve written a lot about magnitude of metric spaces and other structures, but at its most concrete, magnitude is simply an invariant of matrices. A weighting on a matrix is a column vector such that each entry of the column vector is , and it’s an easy lemma that if both and its transpose have at least one weighting then is independent of the weighting chosen. This quantity is the magnitude of . If happens to be invertible then it has a unique weighting, and is simply the sum of all the entries of .
As a trivial example, if is the identity matrix then .
There are a couple of possible meanings for ‘the magnitude of a graph’, because there are (at least!) a couple of ways of turning a graph into a matrix. Not so long ago, I wrote about one of them, in which the matrix encodes the shortest-path metric on the graph. But today, I’ll do something different: when I say ‘the magnitude of ’, I’ll mean the magnitude of the adjacency matrix of . That is, by definition,
for today. (I don’t know whether is always defined, that is, whether always admits a weighting.)
It’s easy to show that magnitude has one of the good properties satisfied by the Lovász number but not the Shannon capacity: it’s multiplicative. That is,
for all graphs and . So if we were to replace maximum diversity by magnitude in the definition
of Shannon capacity, we’d just end up with magnitude again.
How is maximum diversity related to magnitude? The first fact, which I won’t prove here, is as follows: given a similarity matrix ,
the maximum diversity of is the magnitude of some principal submatrix of .
(Recall that a ‘principal submatrix’ of an matrix is one obtained by choosing a subset of , then selecting just those rows and columns indexed by .) In particular, given a graph ,
the maximum diversity of is the magnitude of some induced subgraph of .
(Recall that an ‘induced subgraph’ of a graph is one consisting of a subset of the vertices and all edges between them.) We can take that subgraph to be any independent set of maximal size: for this subgraph has vertices and no nontrivial edges, so by the ‘trivial example’ above, its magnitude is .
Digression If we had to compute the maximum diversity of a similarity matrix, how could we do it? For the present post I don’t think it matters, but the answer’s short enough that I might as well tell you. Given an similarity matrix , let’s say that is ‘good’ if the resulting submatrix admits at least one weighting all of whose entries are nonnegative. Then
This gives an algorithm for computing maximum diversity. It’s slow for large , since we have to run through all subsets of , but at least each step is fast.
In some significant cases, maximum diversity is equal to magnitude. For example, suppose we decide to give two species a similarity of if they’re the same, if they’re different but belong to the same genus, if they’re in different genera but the same family, and so on up the taxonomic hierarchy. Then . The numbers are unimportant here; any other descending sequence would do. In general, given any finite ultrametric space — one satisfying the strengthened triangle inequality
— the matrix with entries has maximum diversity equal to its magnitude.
Let’s have an example: once more, the 5-cycle .
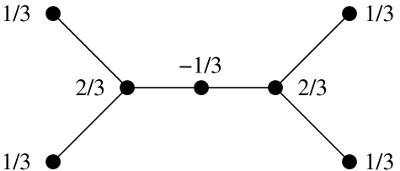
A weighting on the adjacency matrix of a graph is a way of assigning to each vertex a real number , such that for each vertex . There’s a unique weighting on , giving weight to each vertex. The magnitude is the sum of the weights, so .
The maximum diversity is the independence number: .
The Shannon capacity is .
The Lovász number is the same: .
It’s immediately clear that unlike the Lovász number, magnitude is not a helpful upper bound for the Shannon capacity — it’s not an upper bound at all. Indeed, it’s not even an upper bound for the independence number.
So for ,
The equality and the second and third inequalities are true for all . I don’t know whether the first inequality is true in general— that is, whether the magnitude of a graph is always less than or equal to its maximum diversity. Or if not, is it at least true that for all ? Again, I don’t know.
Nor do I know where to go from here. Most of my understanding of magnitude and maximum diversity comes from the context of metric spaces. (For a finite metric space , one uses the similarity matrix given by .) For subspaces of Euclidean space, maximum diversity is always less than or equal to magnitude — which is the opposite way round from what we just saw. That’s because in Euclidean space, passing to a subspace always decreases the magnitude, and by the principle I mentioned earlier, the maximum diversity of a space is equal to the magnitude of some subspace.
Mark Meckes has done a great deal to illuminate the concept of maximum diversity and its relationship to magnitude, putting it into a satisfying analytical setting, not just for finite spaces but also for compact subspaces of Euclidean space (and more generally, so-called spaces of negative type). You can read about this in Positive definite metric spaces and a new paper — Edit: out now! Magnitude, diversity, capacities, and dimensions of metric spaces. But all that’s in a geometrical context quite unlike the graph-theoretic world.
I don’t know whether these ideas lead anywhere interesting. The root of the problem is that I don’t understand much about the magnitude of (the adjacency matrix of) a graph. If you have any insights or observations, I’d very much like to hear them.

Calculating the independence number
Although it doesn’t matter for this post, computing the independence number for an arbitrary “real world” graph in practice is a very interesting problem if you like that kind of thing. It also involves these inequalities, so if you’ll excuse the digression…
Most of the literature on the computing side discusses finding a maximum clique rather than a maximum independent set, so I’ll stick to that to reduce my chances of getting an inequality backwards. We denote this , because it’s the “opposite” of .
The best known way of finding a maximum clique in an arbitrary graph goes something like this: let be some vertex. Then any clique in either does not contain , or contains only and possibly some more vertices adjacent to . We use this to grow a clique, which is conventionally called . Initially is empty. We also have a set of vertices which could potentially be added to be added to . Conventionally this set is called , and initially it contains every vertex in the graph. Now, while isn’t empty, we pick a vertex from . Then we recursively solve the problem with added to and filtered to include only vertices adjacent to (and hence to everything else in ), and then with in neither nor .
Here’s where the inequalities come in. We keep track of the biggest clique found so far, which is usually called the incumbent. At each step in the search, we want a quick way to know whether it’s worth continuing, i.e. whether we might be able to find a clique big enough to unseat the incumbent. One way of doing that is to look at . But isn’t a very good bound.
The best known algorithms right now use a graph colouring as a bound instead: if we can colour the vertices of using colours, cannot contain a clique with greater than k vertices, so if isn’t larger than the size of the incumbent then search at the current point can be abandoned. Calculating the minimum colour number of a graph is also hard, though, but a greedy colouring can be computed very quickly.
There are other considerations (which vertex to select from , which greedy colouring to use, some clever tricks involving bitset encodings, a way of reducing the number of colourings that need to be done, parallelisation, discrepancy based search, …) which aren’t of interest here. But the use of a greedy colouring as a bound works because
The Lovász theta function (of the complement graph) is interesting from an algorithmic perspective because
Also, unlike , can theoretically be computed to within in polynomial time. This would suggest that using a nearest-integer approximation of as a bound would be better than a greedy colouring. Unfortunately calculating the theta function for an arbitrary graph in practice is an open problem…
It doesn’t help that all of these approximations can be arbitrarily bad, and if then there’s no good polynomial time approximation for either or at all.
Still, it’s not all bad news: the smallest graph I’ve found where it’s not possible to compute the clique number exactly has 500 vertices and was designed specifically to screw up these kinds of algorithms. For reasonably sparse graphs we can take it to millions of vertices. So it’s not as horrific in practice as some hard graph problems.