Operads and Phylogenetic Trees
Posted by John Baez
A few years ago, after hearing Susan Holmes speak about the mathematics of phylogenetic trees, I became interested in their connection to algebraic topology. I wrote an article about it:
- John Baez, Operads and the tree of life, Azimuth, 6 July 2011.
In trying to the make the ideas precise I recruited the help of Nina Otter, who was then a graduate student at ETH Zürich. She came to Riverside and we started to work together.
Now Nina Otter is a grad student at Oxford working on mathematical biology with Heather Harrington. I visited her last summer and we made more progress… but then she realized that our paper needed another big theorem, a result relating our topology on the space of phylogenetic trees to the topology described by Susan Holmes and her coauthors here:
- Louis J. Billera, Susan P. Holmes and Karen Vogtmann, Geometry of the space of phylogenetic trees, Advances in Applied Mathematics 27 (2001), 733–767.
It took another half year to finish things up. I could never have done this myself.
But now we’re done! Here’s our paper:
- John Baez and Nina Otter, Operads and phylogenetic trees.
Let me tell you the basic idea….
Trees are important, not only in mathematics, but also biology. The most important is the ‘tree of life’ relating all organisms that have ever lived on Earth. Darwin drew this sketch of it in 1837:
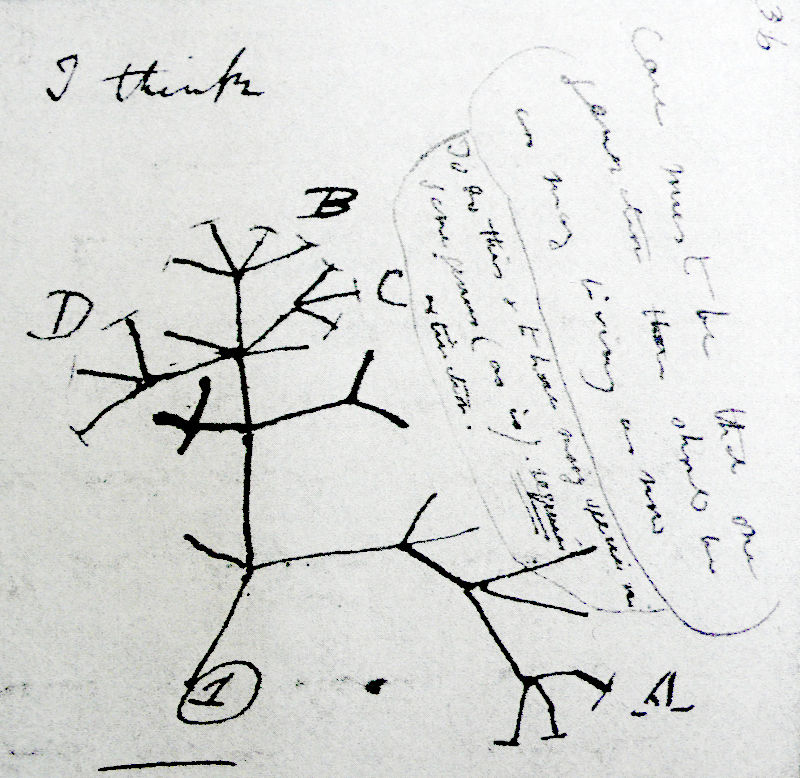
He wrote about it in On the Origin of Species:
The affinities of all the beings of the same class have sometimes been represented by a great tree. I believe this simile largely speaks the truth. The green and budding twigs may represent existing species; and those produced during former years may represent the long succession of extinct species. At each period of growth all the growing twigs have tried to branch out on all sides, and to overtop and kill the surrounding twigs and branches, in the same manner as species and groups of species have at all times overmastered other species in the great battle for life.
Now we know that the tree of life is not really a tree in the mathematical sense. One reason is ‘endosymbiosis’: the incorporation of one organism together with its genetic material into another, as probably happened with the mitochondria in our cells and also the plastids that hold chlorophyll in plants. Another is ‘horizontal gene transfer’: the passing of genetic material from one organism to another, which happens frequently with bacteria. So, the tree of life is really a thicket:
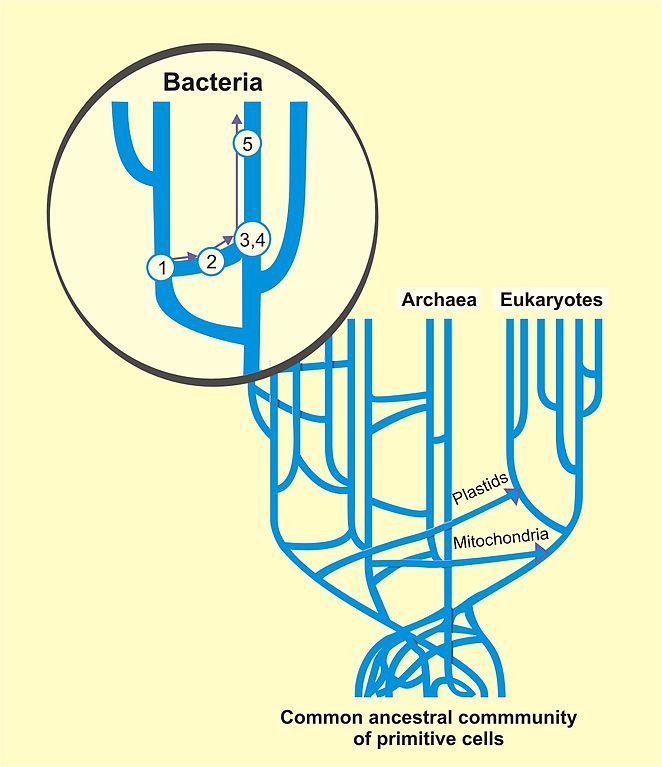
But a tree is often a good approximation, especially for animals and plants in the last few hundred million years. Biologists who try to infer phylogenetic trees from present-day genetic data often use simple models where:
- the genotype of each species follows a random walk, but
- species branch in two at various times.
These are called ‘Markov models’. The simplest Markov model for DNA evolution is the Jukes–Cantor model. Consider one or more pieces of DNA having a total of base pairs. We can think of this as a string of letters chosen from the set {A,T,C,G}:
…ATCGATTGAGCTCTAGCG…
As time passes, the Jukes–Cantor model says the DNA changes randomly, with each base pair having the same constant rate of randomly flipping to any other. So, we get a Markov process on the set
However, a species can also split in two. So, given current-day genetic data from various species, biologists try to infer the most probable tree where, starting from a common ancestor, the DNA in question undergoes a random walk most of the time but branches in two at certain times.
To formalize this, we can define a concept of ‘phylogenetic tree’. Our work is based on the definition of Billera, Holmes and Vogtmann, though we use a slightly different definition, for reasons that will soon become clear. For us, a phylogenetic tree is a rooted tree with leaves labelled by numbers and edges labelled by ‘times’ or, geometrically speaking, ‘lengths’ in . We require that:
- the length of every edge is positive, except perhaps for ‘external edges’: that is, edges incident to the leaves or root;
- there are no 1-ary vertices.
For example, here is a phylogenetic tree with 5 leaves:
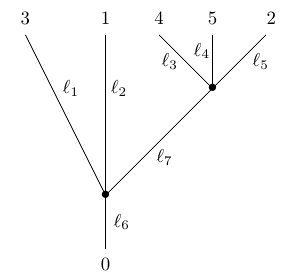
where but we demand that . We draw the vertices as dots. We do not count the leaves and the root as vertices, and we label the root with the number . We cannot collapse edges of length zero that end at leaves, since doing so would eliminate those leaves. Also note that the embedding of the tree in the plane is irrelevant, so this counts as the same phylogenetic tree:
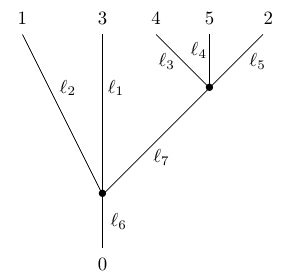
In applications to biology, we are often interested in trees where the total distance from the root to the leaf is the same for every leaf, since all species have evolved for the same time from their common ancestor. These are mathematically interesting as well, because then the distance between any two leaves defines an ultrametric on the set of leaves. However, more general phylogenetic trees are also interesting—and they become essential when we construct an operad whose operations are phylogenetic trees.
Let be the set of phylogenetic trees with leaves. This has a natural topology, which we explain in Section 6 of our paper. For example, here is a continuous path in where we only change the length of one internal edge, reducing it until it becomes zero and we can collapse it:
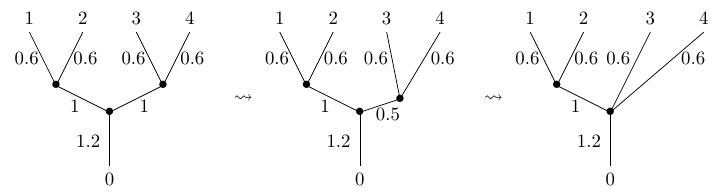
Phylogenetic trees reconstructed by biologists are typically binary. When a phylogenetic tree appears to have higher arity, sometimes we merely lack sufficient data to resolve a higher-arity branching into a number of binary ones. With the topology we are using on , binary trees form an open dense set of , except for . However, trees of higher arity are still important, because paths, paths of paths, etc. in are often forced to pass through trees of higher arity.
Billera, Holmes and Vogtmann focused their attention on the set of phylogenetic trees where lengths of the external edges—edges incident to the root and leaves—are fixed to a constant value. They endow with a metric, which induces a topology on , and there is a homeomorphism
where the data in describe the lengths of the external edges in a general phylogenetic tree.
In algebraic topology, trees are often used to describe the composition of -ary operations. This is formalized in the theory of operads. An ‘operad’ is an algebraic stucture where for each natural number we have a set whose elements are considered as abstract -ary operations, not necessarily operating on anything yet. An element can be depicted as a planar tree with one vertex and labelled leaves:
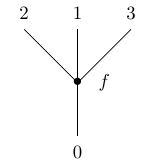
We can compose these operations in a tree-like way to get new operations:
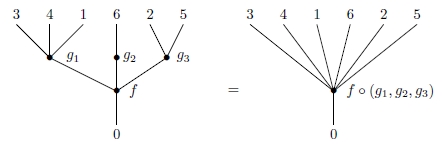
and an associative law holds, making this sort of composite unambiguous:
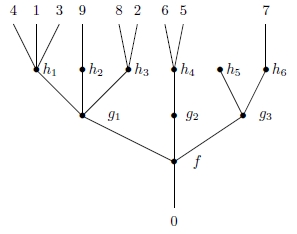
There are various kinds of operads, but in our paper the operads are always ‘unital’, having an operation that acts as an identity for composition. They are also ‘symmetric’, meaning there is an action of the symmetric group on each set , compatible with composition. Further, they are ‘topological’, meaning that each set is a topological space, with composition and permutations acting as continuous maps.
In Section 5 we prove that there is an operad , the ‘phylogenetic operad’, whose space of -ary operations is . This raises a number of questions:
- What is the mathematical nature of this operad?
- How is it related to ‘Markov processes with branching’?
- How is it related to known operads in topology?
Briefly, the answer is that is the coproduct of , the operad for commutative topological semigroups, and , the operad having only unary operations, one for each . The first describes branching, the second describes Markov processes. Moreover, is closely related to the Boardmann–Vogt construction applied to . This is a construction that Boardmann and Vogt applied to another operad in order to obtain an operad whose algebras are loop spaces.
To understand all this in more detail, first recall that the raison d’être of operads is to have ‘algebras’. The most traditional sort of algebra of an operad is a topological space on which each operation acts as a continuous map
obeying some conditions: composition, the identity, and the permutation group actions are preserved, and depends continuously on . The idea is that the abstract operations in are realized as actual operations on the space .
In this paper we instead need algebras of a linear sort. Such an algebra of is a finite-dimensional real vector space on which each operation acts as a multilinear map
obeying the same list of conditions. We can also think of as a linear map
where is the th tensor power of .
We also need ‘coalgebras’ of operads. The point is that while ordinarily one thinks of an operation as having inputs and one output, a phylogenetic tree is better thought of as having one input and outputs. A coalgebra of is a finite-dimensional real vector space on which operation gives a linear map
obeying the same conditions as an algebra, but ‘turned around’.
The main point of this paper is that the phylogenetic operad has interesting coalgebras, which correspond to how phylogenetic trees are actually used to describe branching Markov processes in biology. But to understand this, we need to start by looking at coalgebras of two operads from which the phylogenetic operad is built.
By abuse of notation, we will use as the name for the operad having only unary operations, one for each , with composition of operations given by addition. A coalgebra of is a finite-dimensional real vector space together with for each a linear map
such that:
- for all
- depends continuously on .
Analysts usually call such a thing a ‘continuous one-parameter semigroup’ of operators on .
Given a finite set , a ‘Markov process’ or ‘continuous-time Markov chain’ on is a continuous one-parameter semigroup of operators on such that if is a probability distribution on , so is for all . Equivalently, if we think of as a matrix of real numbers, we demand that its entries be nonnegative and each column sum to 1. Such a matrix is called ‘stochastic’. If is a set of possible sequences of base pairs, a Markov process on describes the random changes of DNA with the passage of time. We shall see that any Markov process on makes into a coalgebra of .
This handles the Markov process aspect of DNA evolution; what about the branching? For this we use , the unique operad with one -ary operation for each > 0. Algebras of are not-necessarily-unital commutative algebras: there is only one way to multiply elements for > 0.
For us what matters most is that coalgebras of are finite-dimensional cocommutative coalgebras, not necessarily with counit. If is a finite set, there is a cocommutative coalgebra whose underlying vector space is . The unique -ary operation of acts as the linear map
where
This map describes the ‘-fold duplication’ of a probability distribution on the set of possible genes. We use this map to say what takes place when a species branches:
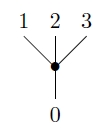
Next, we wish to describe how to combine the operads and to obtain the phylogenetic operad. Any pair of operads and has a coproduct . The definition of coproduct gives an easy way to understand the algebras of . Such an algebra is simply an object that is both an algebra of and an algebra of , with no compatibility conditions imposed.
One can also give an explicit construction of . Briefly, the -ary operations of are certain equivalence classes of trees with leaves labelled and all nodes, except for leaves, labelled by operations in or . While this fact is surely known to experts on operads, it seems hard to find in the literature, so we prove this in Theorem 24. Cat lovers out there will be pleased to note that the proof relies on the existence of an algebraic theory whose algebras are operads.
Given this, it should come as no surprise that the operad is the coproduct . In fact, we shall take this as a definition. Starting from this definition, we work backwards to show that the operations of correspond to phylogenetic trees. We prove this in Theorem 28. The definition of coproduct determines a topology on the spaces , and it is a nontrivial fact that with this topology we have for , where has the topology defined by Billera, Holmes and Vogtmann. We prove this in Theorem 34.
Using the definition of the phylogenetic operad as a coproduct, it is clear that given any Markov process on a finite set , the vector space naturally becomes a coalgebra of this operad. The reason is that, as we have seen, is automatically a coalgebra of , and the Markov process makes it into a coalgebra of . Thus, by the universal property of a coproduct, it becomes a coalgebra of . We prove this in Theorem 36.
Since operads arose in algebraic topology, it is interesting to consider how the phylogenetic operad connects to ideas from that subject. Boardmann and Vogt defined a construction on operads, the ‘ construction’, which when applied to the operad for spaces with an associative multiplication gives an operad for loop spaces. The operad has an interesting relation to . To see this, define addition on in the obvious way, where
Then becomes a commutative topological monoid, so we obtain an operad with only unary operations, one for each , where composition is addition. By abuse of notation, let us call this operad .
Boardmann and Vogt’s construction involves trees with edges having lengths in , but we can equivalently use . Leinster has observed that for any nonsymmetric topological operad , Boardmann and Vogt’s operad is closely related to . Here we make this observation precise in the symmetric case. Operations in are just like phylogenetic trees except that edges may have length . Moreover, for any operad , the operad is a non-unital suboperad of . An operation of lies in if and only if all the external edges of the corresponding tree have length . We prove this in Theorem 40.
Berger and Moerdijk showed that if acts freely on and is well-pointed, then is a cofibrant replacement for . This is true for , the operad whose algebras are topological semigroups. This cofibrancy is why Boardmann and Vogt could use as an operad for loop spaces. But does not act freely on , and is not a cofibrant replacement for . So, it is not an operad for infinite loop spaces.
Nonetheless, the larger operad , a compactification of , is somewhat interesting. The reason is that any Markov process
approaches a limit as . Indeed, extends uniquely to a homomorphism from the topological monoid to . Thus, given a Markov process on a finite set , the vector space naturally becomes a coalgebra of . We prove this in Theorem 38.

Re: Operads and Phylogenetic Trees
This is very cool! It’s neat that “Markov processes with branching” used in genomics have a nice operadic description, and it’s tantalizing that there might be some connection to the original application of operads (iterated loop spaces).
As a step in the latter direction, we might note that if is a -cofibrant replacement for (that is, acts freely on and we have a weak equivalence ), then any -(co)algebra becomes automatically an -(co)algebra. Therefore, the phylogenetic Markov models should also yield -coalgebras, while would be an operad for infinite loop spaces. Then if we could apply some natural contravariant monoidal functor from vector spaces to topological spaces, we would get an actual infinite loop space, and we could ask whether it’s anything familiar to algebraic topologists.